A Fermentation Example for PAT
by Dr. Dieter Bingemann, Cicely Rathmell
Published 2/21/20
The growing use of cellular and biomolecular processes in energy, medical, and environmental applications has created a need for real-time sensing of both reactants and products through process analytical technology (PAT). Raman is already an established process monitoring tool used in PAT, providing excellent analytical selectivity while being particularly insensitive to water present as solvent or moisture within the sample.
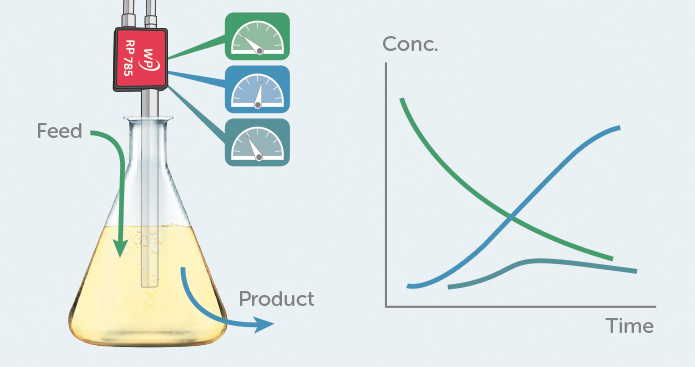
In this application note, we explore the use of Raman to monitor a simple bioprocess, fermentation of glucose (a common feedstock) with yeast, a microorganism often used in biotechnology. Using a probe-based WP 785 Raman system, we calibrated for glucose concentration in a yeast solution, then monitored the glucose content throughout fermentation. Applying multivariate tools to the Raman spectra also allowed us to clearly follow the overall reaction progress and identify the main reactants: glucose, ethanol, and carbon dioxide. The results show how a sensitive, stable Raman system can be successfully applied in PAT for bioprocessing applications, even when fluorescence background is present.
Raman, a PAT solution for Biotechnology
Microbial biotechnology harnesses the fermentation process of microorganisms to generate value-added products such as biogas/bioethanol for energy, processed food products, recombinant vaccines/antibiotics, and key chemicals such as enzymes, organic acids, alcohols, and polymers. It can also be put to environmental use in wastewater treatment for the removal of toxic materials. materials. Microorganisms can quickly be cultured in large quantities, and their enzyme production enhanced through genetic manipulations to further improve and target these biocatalysts.
Process analytical technology, and more specifically Raman spectroscopy, can be used to provide real-time process monitoring of fermentation. Raman’s ability to distinguish many different chemicals in parallel, its insensitivity to water, and the ability to deploy using a simple probe-based system makes it a well-suited tool to monitor and control biotechnology processes with greater consistency and yield.
To demonstrate, we applied Raman spectroscopy to a representative biotechnology process, the fermentation of glucose with yeast, using a 785 nm Raman spectrometer connected to a dip probe inserted into the fermentation mixture. We analyzed the resulting set of Raman spectra both quantitatively, by extracting the glucose concentration and following the overall reaction progress, as well as qualitatively, by identifying the main reactants of the process through their Raman spectra.
Probe-based Raman Process Monitoring
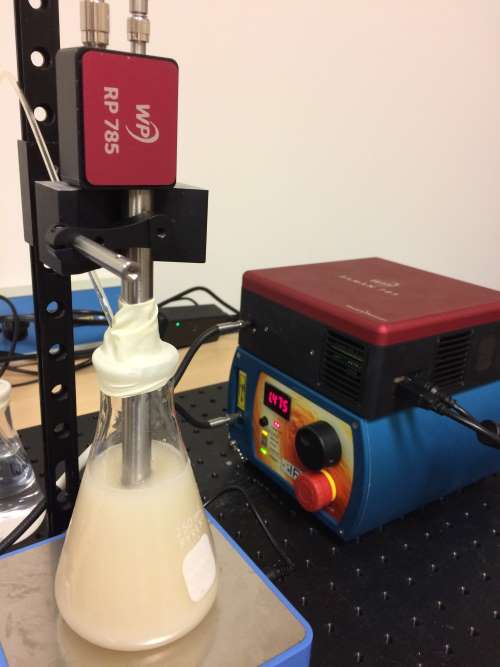

The Raman process monitoring system was based upon a WP 785 spectrometer covering 270-2000 cm-1 with 10 cm-1 resolution, equipped with a TEC-regulated detector (figure 2). An external 785 nm laser provided 350 mW of excitation light, fiber-coupled into our RP 785-VAR probe fitted with a special process barrel. This unique probe design features an interchangeable tip to allow probe reconfiguration for different sample types and working distances. In this case, it was configured as a 15 cm long immersion probe. The Raman emission collected by the probe was transferred to the spectrometer with a 0.39 NA fiber to fill the f/1.3 input aperture of the spectrometer fully for greatest sensitivity.
The Raman spectra for both calibration and monitoring were collected with an integration time of 5 seconds. Calibration spectra were recorded 20 times for each solution. During fermentation, 20 spectra were collected and averaged, repeated at 120 second intervals. All spectra were saved with a “laser-on dark” blank subtracted. This blank measurement was performed using the same sample container filled with water.
A total of 30 calibration solutions were used to establish the glucose concentration model, prepared from stock glucose and yeast solutions to represent all possible permutations of glucose concentration (0-50 g/L) and yeast (0-10 g/L), as shown in Table 1. The glucose solutions were allowed to equilibrate via mutarotation prior to the calibration measurements.
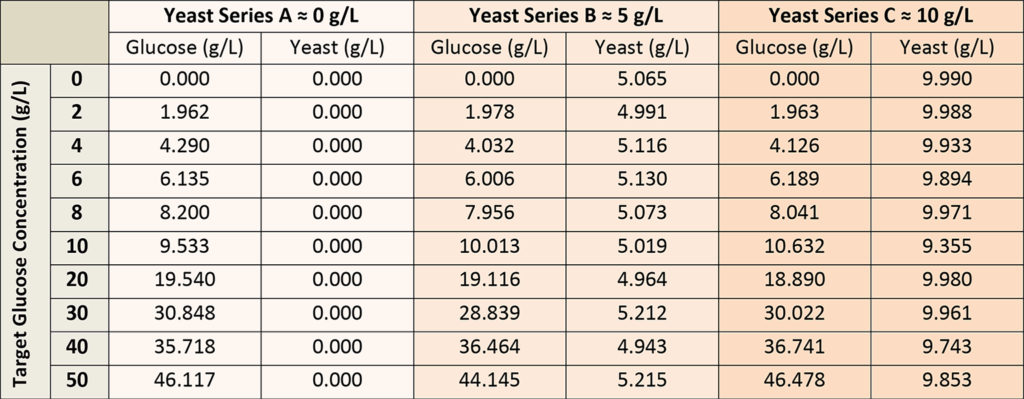
It was important to include larger concentrations of yeast in the calibration solutions, as yeast contributes fluorescence background to the Raman spectra. Each calibration solution was measured in a large round sample vial, with the dip probe penetrating about 5 mm into the solution. The vial was held over a black, empty void to avoid any back reflection from the mount, aside from the vial bottom. In the case of calibration solutions with yeast, the calibration solutions were stirred to reduce signal fluctuations caused by the yeast.
The fermentation experiment was performed in a 250 mL Erlenmeyer flask. The solution was stirred continuously with a magnetic stirrer, with the dip probe penetrating well past the (small) vortex surface into the solution. The flask was sealed with a rubber membrane across the top; the dip probe and a piece of tubing penetrated the membrane. The other end of the tubing was dipped into a second flask filled with water, allowing for pressure equilibration while maintaining anaerobic conditions for the fermentation.
The fermentation solution was prepared with warm water to speed the reaction, with initial concentrations of approximately 50-60 g/L glucose and 2-3 g/L yeast. The fermentation was observed over a 14-hour period at 120 second intervals.
Glucose Calibration – Knowing our Sugars
The unprocessed Raman spectra for all glucose and yeast concentrations are shown in figure 3a. Here we see three groups of spectra, corresponding to the three yeast series: 0, 5, and 10 g/L. As yeast concentration increases, so does the fluorescence background, due to its complex organic structure. Within each yeast series, we see spectral variation corresponding to the concentration of glucose; the higher concentrations show increasingly larger peaks indicative of the glucose structure, particularly around 500 and 1100 cm-1.
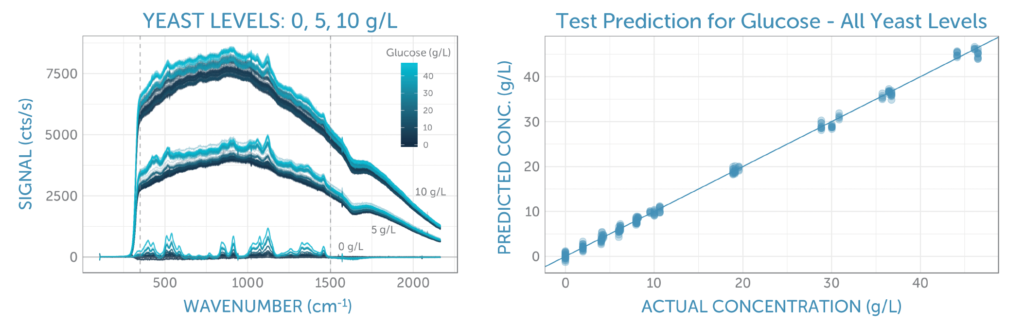
To develop a predictive model for glucose, we correlated the middle portion of the Raman spectra, as indicated by the two vertical dashed lines in figure 3a, with the glucose concentration using PLS (partial least squares) regression. In doing so, we randomly split the repeat spectra recorded for each calibration solution into equally sized training and test sets. The training set was used to optimize the calibration method through 5-fold cross-validation, and the test set was reserved to determine the final prediction error.
Three pre-processing approaches were tested: raw (no pre-processing), baseline subtraction using ALS (asymmetric least squares with lambda = 4 and p = 0.001), and first derivative smoothing using a Savitzky-Golay filter (in second order over a 9-pixel window). While the latter two methods were effective at removing the yeast fluorescence background, we found surprisingly that analyzing the unprocessed spectra yielded the lowest cross-validation error for glucose prediction, closely followed by regression of the spectra after baseline subtraction. The final calibration regression applied to the test set, as shown in figure 3b, yielded a prediction error (RMSEP) of about 0.7 g/L, independent of the yeast concentration. With this, we could be confident in our prediction model regardless of yeast concentration and ensuing fluorescence background.
Real-time Raman Process Monitoring
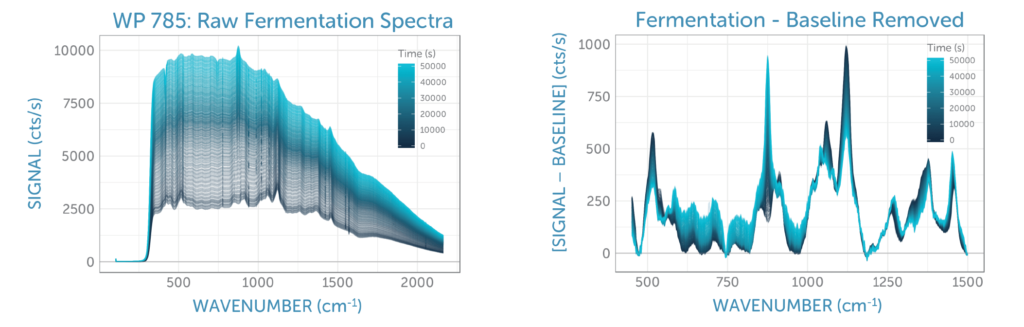
We continuously monitored the Raman spectra of the fermentation sample for 14 hours, taking spectra every 120 seconds. The sequence of unprocessed spectra plotted in figure 4a show a transformation of spectral features over time as glucose was consumed by the yeast and ethanol was generated. The relative changes and the overall signal quality are more clearly visible after removal of the yeast fluorescence background using ALS (asymmetric least squares), as shown in figure 4b.
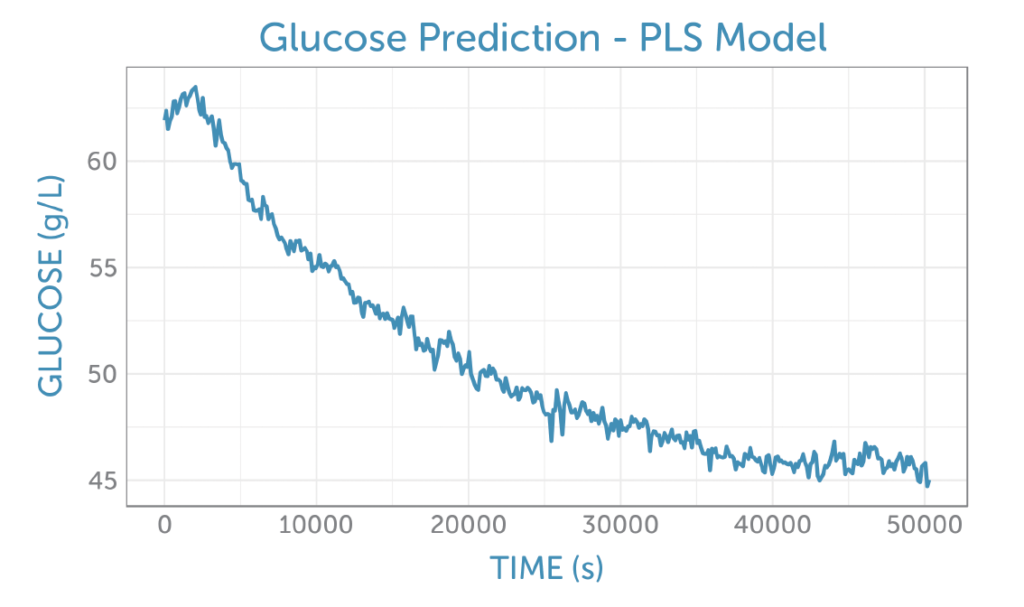
Applying the PLS glucose calibration model to the unprocessed fermentation spectra as they were captured, we were able predict the glucose concentration in real-time during the fermentation, as shown in figure 5. No offline reference measurement was available for validation of the accuracy, but the prediction shows a low noise level for the concentration measurement.
PCA – A More Holistic View
Curious to explore what other process monitoring information could be extracted from the spectra over time, we performed principal component analysis (PCA) on the entire spectral data set acquired during the fermentation. As we were interested in a chemical interpretation of the process, we chose not to center the PCA so that the resulting scores and loadings would be more reminiscent of concentrations and component spectra rather than drawing out the harder to interpret relative spectral changes. In addition, to ignore the dominant yeast contribution to the signal, we performed PCA on the baseline-corrected spectra shown in figure 4b.
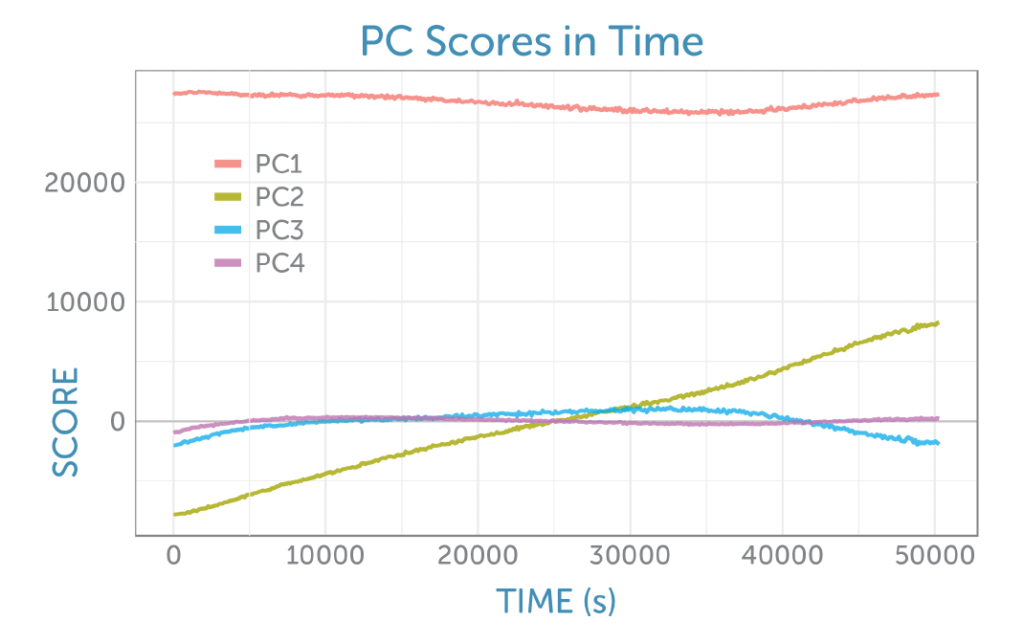
Four principal components were identified, each rich in information about the reactants and products. The PCA scores as a function of time are shown in figure 6.
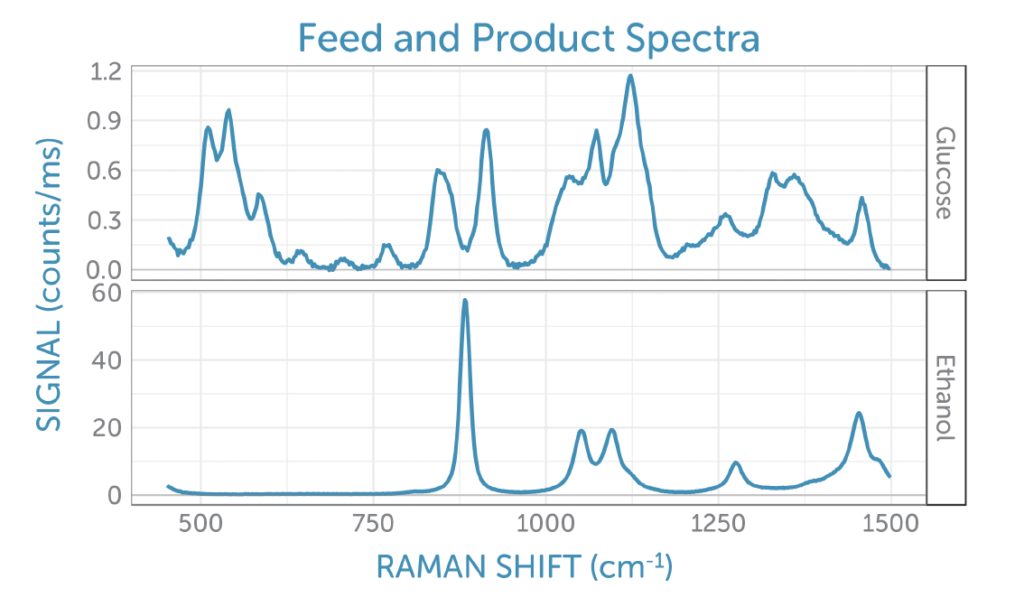
To better interpret the scores, we can look at the corresponding loadings plotted as Raman spectra on the corresponding frequency axis. Unlike actual Raman spectra, loadings can contain both positive and negative contributions and often have the appearance of difference spectra. These are easier to interpret when compared to the reference spectra taken for glucose (the feedstock, which is consumed during fermentation) and ethanol (the product generated), shown in figure 7.
The loadings for the four principal components are shown in figure 8. The loading for the first PCA component resembles the glucose Raman spectrum, while the second component has the appearance of the difference spectrum between the reactant glucose and the main product ethanol as shown by PC2 in figure 6. This second component’s scores continuously rise during the fermentation, providing a visual indicator of the overall progress of the reaction through the ratio of these two components, which is ideal for process monitoring.
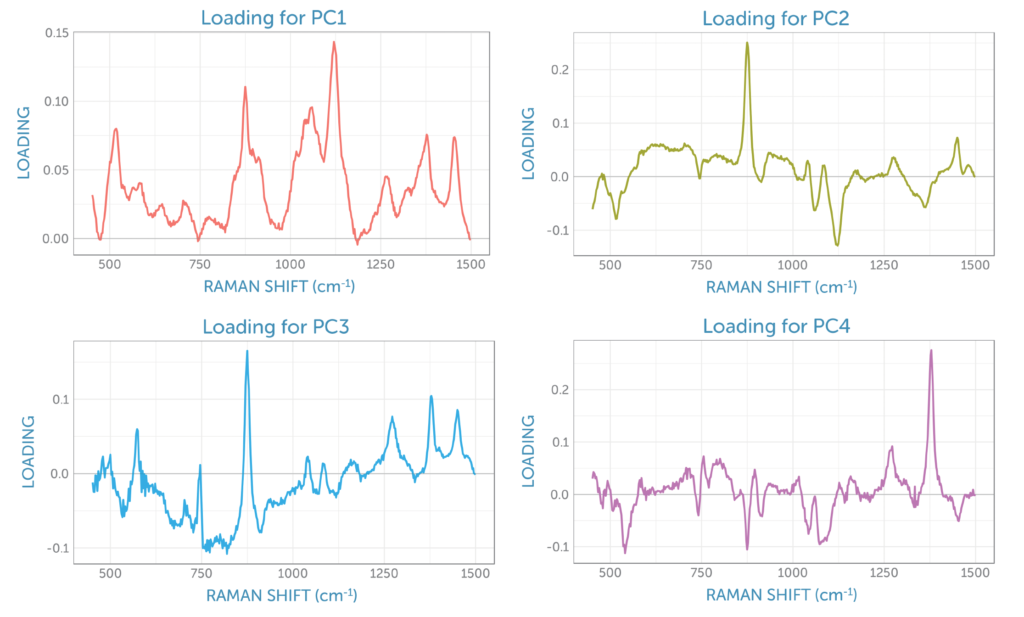
The third loading contains spectral signatures from four different contributions: the reactant glucose, the product ethanol, the dissolved by-product carbon dioxide with its peaks at 1280 and 1382 cm-1, and two residual peaks from sapphire (the dip probe optical material), with peaks at around 580 and 750 cm-1. The fourth loading appears to be dominated by carbon dioxide, which remains relatively consistent in solution throughout fermentation due to the stirring.
Raman Process Monitoring Success
Using a modular 785 nm Raman system fitted with an immersion probe, we continuously recorded spectra during a 14-hour fermentation of glucose with yeast. Using a PLS regression model for the glucose concentration in the presence of yeast we quantitatively monitored the decrease in glucose during fermentation with high signal to noise.
Additional principal component analysis and plotting of the time-dependent scores found that the second component captured the reaction progress of the transformation of glucose into ethanol. Additional reaction products, such as dissolved carbon dioxide, could be identified in the higher components’ loadings.
The results of this fermentation experiment show that, even with the presence of significant fluorescence background, Raman spectra can provide detailed information about the progress of biotechnological processes. Other processes that are well-matched to the strengths of Raman spectroscopy could also be monitored using a similar approach, from solvent removal to crystallizations, including the identification of polymorphs. With a sensitive, portable system like the WP 785, Raman process monitoring of fermentation is just the beginning.


